by Gholamhossein Amini Chermahini, MD, Columbus, Ohio
Editor’s Note: Dr. Amini Chermahini and his colleagues were awarded the Best Poster Prize at this year’s FSHD International Research Congress. Here he explains the significance of this work.
The FSHD field has made tremendous progress during the last decade or so, beginning with the identification of the DUX4 gene as a root cause of the disease. Several new experimental FSHD therapies are emerging, and most are designed to turn DUX4 “off” or reduce its levels.
As these new treatments reach human clinical trials, it becomes important for clinicians to have “outcome measures” that can be used to determine if a treatment is working or not. One such outcome measure could detect whether a drug has reduced the amount of DUX4 present in a treated muscle compared to an untreated muscle. Unfortunately, because DUX4 is only active in one out of a thousand muscle nuclei at a given time, it is like searching for the proverbial needle in a haystack. It is difficult to find and detect, and to our knowledge there were no established methods to reliably measure DUX4 in human muscles.
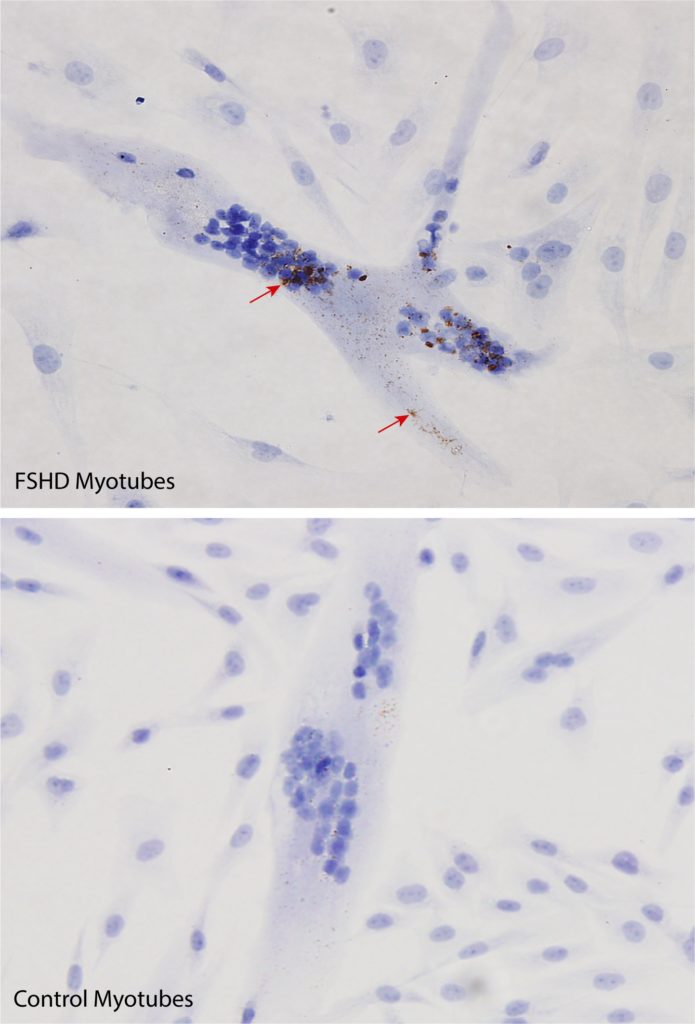
The goal of our study was to test a new and highly sensitive method to detect DUX4. In our RNAscope assay, an RNA in situ hybridization (ISH) technology, several probes* target the specific area in tandem on the mRNA (messenger RNA),** which increases the specificity of DUX4 detection. At the time we began this study, the vendor had not developed DUX4-targeted RNAscope probes and we therefore initiated our study using custom-designed DUX4 RNAscope probes. We found that this method was capable of detecting very low levels of DUX4 in isolated FSHD muscle cells grown in the lab (Fig 1). Importantly, we also found that this approach could measure reductions in DUX4 after FSHD muscle cells were exposed to an experimental therapy.
This study is published now in the RNA journal (2019) titled “RNAscope in situ hybridization-based method for detecting DUX4 RNA expression in vitro” and also was presented at the IRC 2019 in Marseille. We were honored to receive the “Best Poster Prize,” which is very encouraging and at the same time testifies to the importance of the subject in the field. The next step of this study is to determine if the method will work in muscle biopsies donated by FSHD patients. If so, we propose that it may be useful as an outcome measure in future clinical trials.
*Probe. A fragment of RNA or DNA that recognizes part of the genetic sequence of DUX4 and binds to it, like a key fitting a lock. A radioactive or fluorescent tracer on the probe then enables the probe to be detected.
**mRNA. Genetic information flows from DNA to RNA to protein. The first step employs a process called transcription to read a strand of DNA and synthesize a corresponding piece of messenger RNA (mRNA); the second step employs a process called translation to read that piece of mRNA and build a corresponding protein.


Leave a Reply